Biochemical investigation of monoclonal gammopathies
Monoclonal gammopathy (MG) refers to the presence of monoclonal immunoglobulin produced by clonally expanded plasma cells or immunoglobulin-expressing lymphocytes. MG is a key feature of a wide spectrum of diseases ranging from the indolent MG of undetermined significance to the overt multiple myeloma. In this article, we discuss the utility and pitfalls of common biochemical techniques used to detect MG.
by Dr Michelle L. Parker and Dr Pak Cheung Chan
Introduction
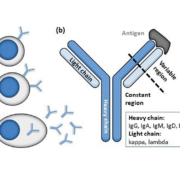
The monoclonal immunoglobulins or ‘M-proteins’ detected in monoclonal gammopathy (MG) are produced by clonally expanded plasma cells, or less frequently by immunoglobulin-expressing lymphocytes at different stages of maturation. The prevalence of MG in the general population over 50 years of age is approximately 3 % and increases with age. M-proteins secreted by plasma cells (Fig. 1a) can be partial or intact immunoglobulins, with the latter consisting of two heavy chains and two light chains that together form a Y-shaped structure with constant and highly variable antigen-binding domains (Fig. 1b). M-proteins that are immunologically functional may cause disease by directly binding to self-antigens, e.g. in some peripheral neuropathy. Other unique chemical properties may cause the M-protein to transform into insoluble amyloids, to increase plasma viscosity, or even to block capillary blood flow by precipitating out at the low temperatures in the extremities. As the production of M-protein increases, the mass effect can be exerted through the expanded clonal plasma cells compressing neighbouring cell lineages in the bone marrow, resulting in reduced red blood cell production (anemia), pan-leukopenia (recurrent infections), thrombocytopenia (bleeding diathesis), suppressed non-involved plasma cells (immune paresis) and bone resorption (hypercalcemia and bone lesions). Large amounts of circulating M-protein could promote plasma hyperviscosity, thrombosis, and tissue and organ damage. For example, excess filtered free light chains in multiple myeloma can directly damage the kidney proximal tubules, form amyloids rupturing glomeruli and form obstructive casts in the distal tubules leading to cell death and nephritis. In general, measured M-protein concentration is taken to reflect the tumour burden and is prognostic for disease progression or survival, e.g. in monoclonal gammopathy of undetermined significance (MGUS), smouldering myeloma and multiple myeloma.
Conditions associated with MG cover a wide range of clinical presentations and severity, including MGUS, multiple myeloma, P.O.E.M.S., light chain deposition disease, plasmacytoma, Waldenstrom’s macroglobulinemia, non-Hodgkin’s lymphoma and chronic lymphocytic leukemia. In some of these diseases, the severity of tissue or organ damage may not be related to the M-protein concentration. For example, in some amyloid light chain (AL)-amyloidosis, extensive kidney damage is reflected by massive proteinuria, yet the circulating monoclonal free light chain can be barely, or not at all, demonstrable by serum and/or urine testing [1]. Nevertheless, the presence of an M-protein can be a defining hallmark of many of these conditions and its detection provides a critical link to their final diagnosis.
Biochemical detection of monoclonal immunoglobulins
Five common biochemistry tests form the core of first-line MG investigations and will be discussed below: serum protein electrophoresis (SPE), serum immunofixation electrophoresis (IFE), urine protein electrophoresis (UPE), urine immunofixation electrophoresis (uIFE), and serum free light chain (sFLC) assays. Other techniques such as mass spectrometry-based assays and HevyliteTM analysis are increasingly available for specific circumstances but will not be discussed here.
SPE and UPE
SPE and UPE resolve serum and urine proteins respectively into five or six major fractions, viz. albumin, alpha-1, alpha-2, beta (total beta, or beta-1 and beta-2 depending on resolution), and gamma (Fig. 2). If a monoclonal antibody is present, an additional peak may be observed, most frequently in the gamma (hence the term gammopathy) (Fig. 2) but other regions such as beta and alpha-2 are also possible. Estimating the size of this extra peak gives the amount of M-protein present and is one of the recommended methods for monitoring disease activity. However, the detection of M-protein this way requires that it is readily distinguished from background polyclonal immunoglobulins or other co-migrating proteins, which not only limits the analytical sensitivity to around 0.5–2.0 g/L [2] and prevents its use to rule-out low abundance M-proteins [3, 11], but also limits the accuracy of quantification especially at low M-protein concentrations and/or high background in any electrophoretic regions.
Importantly, an ‘abnormal’ peak identified by SPE does not prove that it is an endogenous monoclonal immunoglobulin, as the peak may be due to a haptoglobin variant, iodinated contrast material, aminoglycoside, administered biologics, or increases in other proteins such as tumour markers, transferrin in severe iron deficiency, C-reactive protein in acute inflammation, and fibrinogen in plasma or incompletely clotted serum [4]. Similarly, a positive finding in UPE can only be regarded as presumptive and should be confirmed by techniques such as IFE.
Historically, qualitative deviations from the expected SPE pattern have been taken to imply clinical conditions such as bisalbuminemia, acute-phase inflammatory response, alpha-1-antitrypsin deficiency, nephrotic syndrome, cirrhosis, hypogammaglobulinemia, etc. However, not all of these conditions as predicted by SPE patterns have been validated, nor have their clinical utility in terms of MG investigation been established [5].
IFE and uIFE
For IFE, a combination of antisera against the heavy chains (IgG, IgA, IgM, IgD, IgE), the two light chains (total kappa and total lambda) and/or the free light chains (free kappa and free lambda) is selected and separately overlaid on the electrophoresed sample. Immuno-precipitation results in a blush of staining in the presence of polyclonal immunoglobulins, while a discrete band indicates the presence of an M-protein and its isotype is determined when discrete bands in the heavy and light chain lanes are aligned (Fig. 2 inset). This immunological detection not only characterizes the M-protein whose isotype provides prognostic information, but also improves the analytical sensitivity (typically 0.2 to 0.5 g/L) enabling detection of M-proteins even when the SPE pattern is visibly normal [2]. However, a notable short fall is that the interpretation is unavoidably subjective especially when bands are faint or not well defined.
In uIFE, the focus is to detect monoclonal free light chains or Bence Jones proteins that passed through the kidneys unabsorbed. In normal individuals, immunoglobulin light chains are produced in slight excess of the heavy chains and are secreted into the circulation. Because of their small sizes, free light chains are readily filtered through the glomeruli but are efficiently absorbed in the proximal tubules. Thus, in patients with MG, the detection of monoclonal free light chains in urine usually indicates an increased production exceeding renal reabsorbing capacity, compromised reabsorption, or both. Since the secretion of free light chains into the circulation is sporadic throughout the day, a ‘pooled’ sample such as a 24-h urine collection usually improves the sensitivity as well as the reliability of urine testing, although a first-morning urine has also been accepted for initial investigations.
sFLC assays
The fully automated sFLC measures polyclonal immunoglobulin free light chains individually with high analytical sensitivity (down to mg/L) and targets the light chain epitopes that are otherwise hidden when bound to heavy chains (Fig. 3)[2]. Patients with MG often have increased concentrations of the involved free light chains, resulting in a skewed free kappa/lambda ratio as the uninvolved free light chains remains normal or suppressed. A skewed ratio not only supports the diagnosis of MG but also provides prognostication information on malignant progression for MGUS, smouldering myeloma and multiple myeloma. A free kappa/lambda ratio >100 has even been taken as a defining feature for multiple myeloma [6].
Similar to many other immunoassays, the sFLC assay is subject to antigen excess and displays dilutional non-linearity, raising concern over the accuracy of results at both high concentrations (variation due to different dilution response) and low concentrations (high dose hook effect). Additionally, falsely abnormal free kappa/lambda ratios have been reported in individuals with polyclonal gammopathy, hospitalized patients and patients with renal dysfunction. In one study, the reported positive predictive value of an abnormal ratio amongst primary care patients was only 39 % [7], underscoring the high false-positive rate in unselected patients. Although there are sFLC assays reportedly less susceptible to these limitations [8], a general lack of standardization renders the results non-commutable and values cannot be interchanged between methods.
Diagnostic testing algorithms
Although the biochemical tests discussed above play an important role in the detection of M-proteins, the information that each test provides does overlap substantially, and different test combinations may be required for different monoclonal gammopathies. Moreover, these tests tend to be costly, labour intensive, and/or require expertise for result interpretation. There is ongoing debate on the optimal testing algorithm due to competing priorities such as maximizing clinical sensitivity or diagnostic efficiency, streamlining workflows, improving economic feasibility, and reducing unnecessary or redundant testing.
With a primary goal of maximizing clinical sensitivity, the International Myeloma Working Group (IMWG) recommends SPE, IFE and sFLC as first-line tests for confirming multiple myeloma and other plasma cell disorders, with the addition of 24-hour urine studies only if AL-amyloidosis is suspected [2, 8]. Although the recommendation falls short of indicating that these tests may be performed in tandem depending on findings, it does represent a welcomed change to previous versions as 24-h urine samples are inconvenient to collect and UPE and uIFE are expensive to perform. Although sFLC testing has largely obviated the need for first-line urine studies, no single serum test has adequate clinical sensitivity for screening all plasma cell disorders [8, 9]; in one large study, SPE, IFE and sFLC had clinical sensitivities of just 79, 87 and 74 % respectively [3].
The optimal combination of first-line and reflexed tests remains difficult to determine owing to the wide spectrum of MG diseases. There is substantial redundancy if SPE and IFE are performed simultaneously. IFE contains a protein lane that provides the same qualitative detection of M-proteins as SPE. A separate SPE only provides additional information regarding quantity of the M-protein, as there are no true positives that would be missed by IFE but identified by SPE. For economic and other reasons, SPE is often performed initially and is reflexed to IFE for confirmation if SPE presents with features suggestive of an M-protein, including the observation of restricted staining or a clearly discrete band, increased beta fraction [10], or decreased gamma fraction [11]. However, this approach has been shown to miss up to 20 % of cases [3, 10–12] as some M-proteins, especially free light chains and those existing in small concentrations, may not present with any abnormal features in SPE. Recently, it was argued that the increased sensitivity of IFE over SPE warrants its use as the first-line screening test, despite being more expensive and labour intensive. The use of modified IFE protocols such as combined light chain immunofixation (a mixture of anti kappa and anti-lambda antisera), or the penta-IFE using a mixture of five antisera (anti IgG, IgA, IgM, kappa, and lambda) seems to make this approach more feasible. The counterpoint to this approach, though, is that the detection of very low concentration M proteins by IFE may lead to unnecessary investigation of transient or low risk conditions [13]. On the other hand, without full characterization of the M-protein (both isotype and concentration), it may be premature to judge the significance of an M-protein based only on its low concentration.
Concluding Remarks
Clearly, further studies are needed to balance the competing priorities of various testing algorithms and provide evidence-based approaches to MG investigations suited to the diverse clinical environments, ranging from family practice to speciality hematology clinics. Irrespective of the algorithm used, it is good practice to interpret laboratory findings within the specific clinical context to mitigate the risk of false-positive or false-negative test results.
References
1. Truong D, Blasutig IM, Kulasingam V, Chan PC. A patient with monoclonal gammopathy-related nephrotic syndrome revealed no electrophoretic “nephrotic pattern” or skewed free light chain ratio. Clin Biochem 2018; 51: 110–111.
2. Dispenzieri A, Kyle R, Merlini G, Miguel JS, Ludwig H, Hajek R, et al. International Myeloma Working Group guidelines for serum-free light chain analysis in multiple myeloma and related disorders. Leukemia 2009; 23(2): 215–224.
3. Katzmann JA, Kyle RA, Benson J, Larson DR, Snyder MR, Lust JA, et al. Screening panels for detection of monoclonal gammopathies. Clin Chem 2009; 55(8): 1517–1522.
4. McCudden CR, Jacobs JFM, Keren D, Caillon H, Dejoie T, Andersen K. Recognition and management of common, rare, and novel serum protein electrophoresis and immunofixation interferences. Clin Biochem 2018; 51: 72–79.
5. Chan PC, Chen Y, Randell EW. On the path to evidence-based reporting of serum protein electrophoresis patterns in the absence of a discernible monoclonal protein – A critical review of literature and practice suggestions. Clin Biochem 2018; 51: 29–37.
6. Rajkumar SV, Dimopoulos MA, Palumbo A, Blade J, Merlini G, Mateos MV, et al. International Myeloma Working Group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol 2014; 15(12): e538–548.
7. Hill PG, Forsyth JM, Rai B, Mayne S. Serum free light chains: An alternative to the urine Bence Jones proteins screening test for monoclonal gammopathies. Clin Chem 2006; 52(9): 1743–1748.
8. Tate JR, Graziani MS, Mollee P, Merlini G. Protein electrophoresis and serum free light chains in the diagnosis and monitoring of plasma cell disorders: laboratory testing and current controversies. Clin Chem Lab Med 2016; 54(6): 899–905.
9. Willrich MAV, Murray DL, Kyle RA. Laboratory testing for monoclonal gammopathies: focus on monoclonal gammopathy of undetermined significance and smoldering multiple myeloma. Clin Biochem 2018; 51: 38–47.
10. Chan PC, Lem-Ragosnig B, Chen J. Diagnostic implications of enumerating and reporting beta fraction(s) for the detection of beta-migrating monoclonal immunoglobulins in serum protein electrophoresis. Clin Biochem 2018; 53: 77–80.
11. Chan PC, Chen J. Value of reflex testing based on hypogammaglobulinemia as demonstrated in serum protein electrophoresis. Clin Biochem 2015; 48: 674–678.
12. Pretorius CJ. Screening immunofixation should replace protein electrophoresis as the initial investigation of monoclonal gammopathy: Point. Clin Chem Lab Med 2016; 54(6): 963–966.
13. Smith JD, Raines G, Schneider HG. Should routine laboratories stop doing screening serum protein electrophoresis and replace it with screening immune-fixation electrophoresis? No quick fixes: Counterpoint. Clin Chem Lab Med 2016; 54(6): 967–971.
The authors
Michelle L. Parker1 PhD, Pak Cheung Chan*1,2 PhD, DABCC, FCACB
1Department of Laboratory Medicine and Pathobiology, University of Toronto, Toronto, ON, Canada
2Department of Laboratory Medicine & Molecular Diagnostics, Sunnybrook Health Sciences Centre, Toronto, ON, Canada
*Corresponding author
E-mail: pc.chan@sunnybrook.ca